class: center, middle, inverse, title-slide .title[ # Conda and Herper<br /> <html><br /> <br /> <hr color='#EB811B' size=1px width=796px><br /> </html> ] .author[ ### Rockefeller University, Bioinformatics Resource Centre ] .date[ ### <a href="https://rockefelleruniversity.github.io/RU_reproducibleR/" class="uri">https://rockefelleruniversity.github.io/RU_reproducibleR/</a> ] --- ## What is Conda? [Conda](https://conda.org/) is an open-source and cross-platform package and environment management system. It was originally developed to help manage packages for Python (similar to Renv), but that has expanded beyond python and is very popular with R users as well. --- ## How does Conda work? 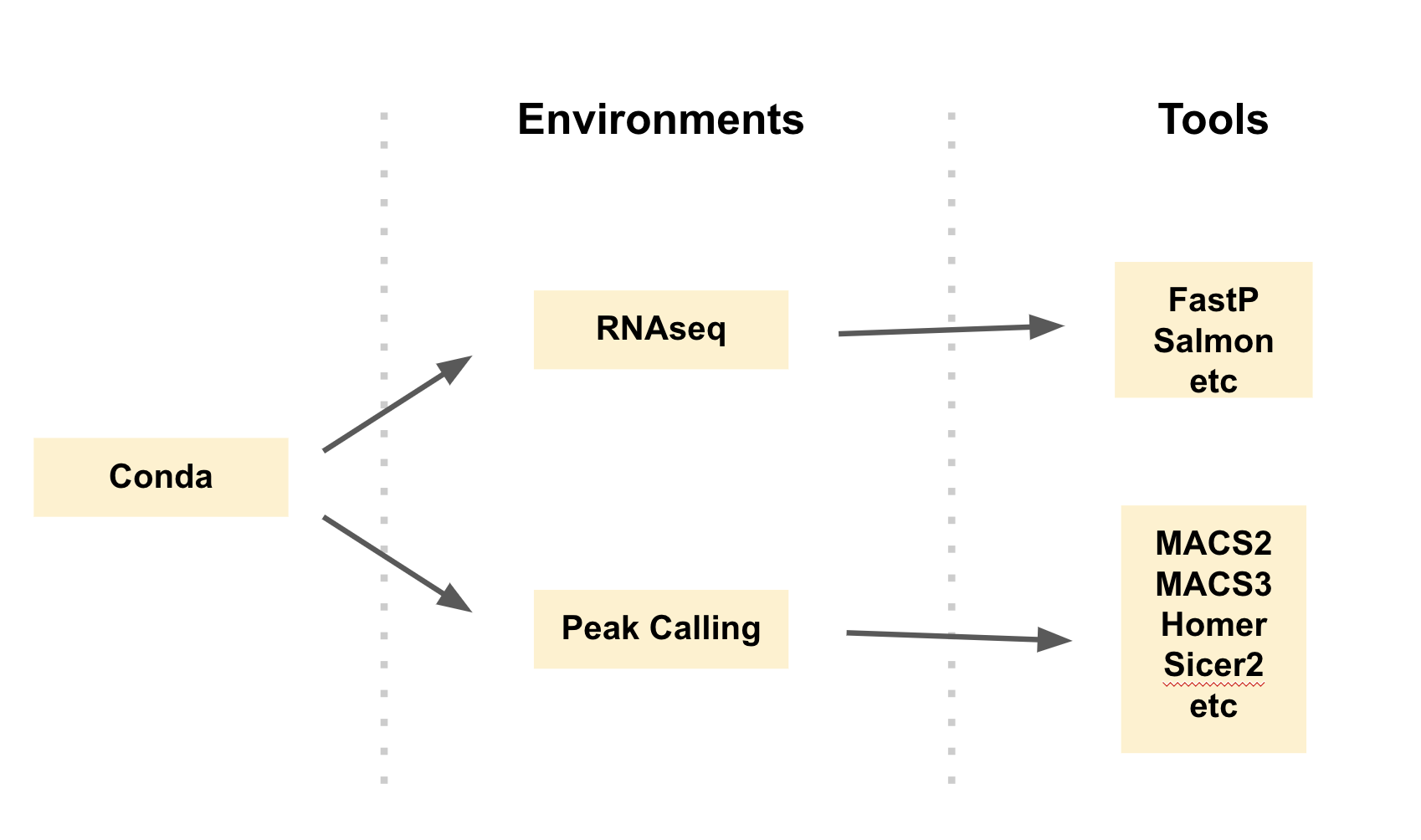 --- ## Why Conda? * Simplify Package Management * Cross-Platform Compatibility * Extensive Package Ecosystem * Environment Management --- ## Conda vs miniconda vs ... There are many different repository tools in the Conda ecosystem. They all have slight differences. Conda - The core package manager Miniconda - A minimal install of Conda Anaconda - A maximal install of Conda Mamba - A reimplementation of Conda built in C++ [FASTER] Minimamba - A minimal install of Mamba --- ## Conda vs miniconda vs ... 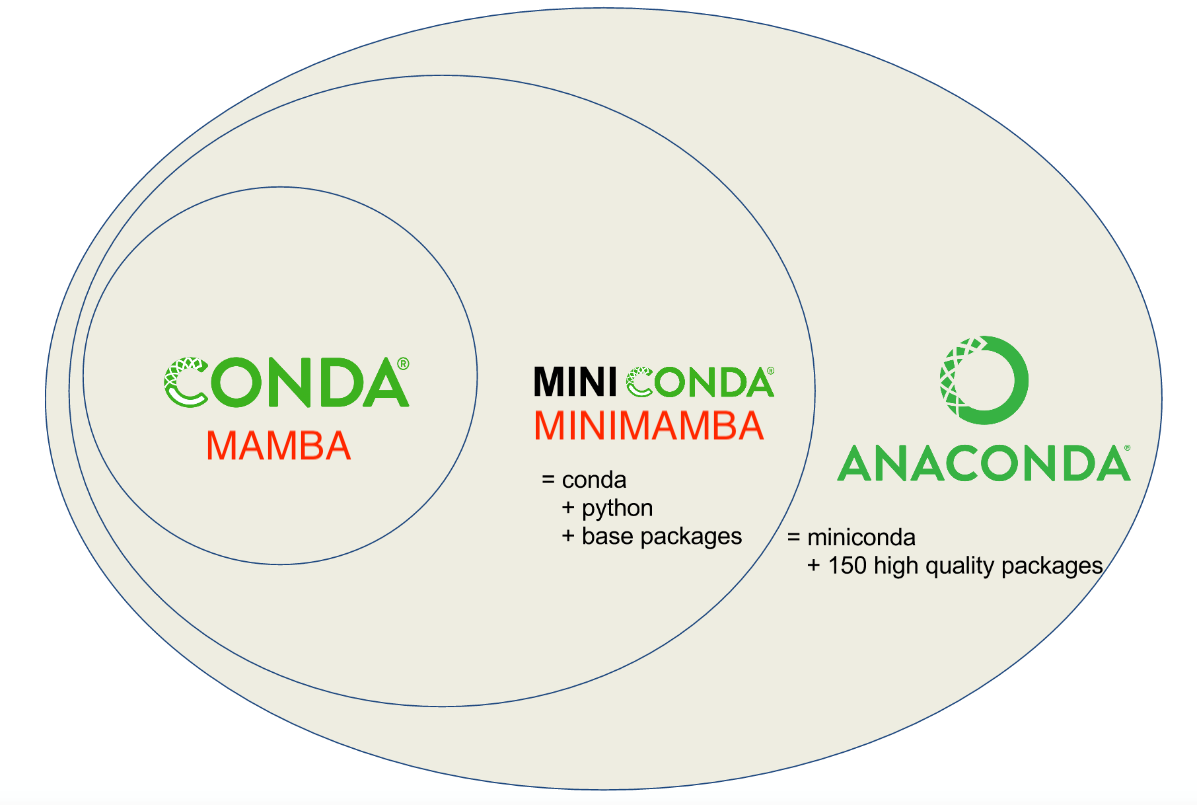 [Python Envs Guide](https://www.whiteboxml.com/blog/the-definitive-guide-to-python-virtual-environments-with-conda) ] --- ## Conda vs miniconda vs ... We recommend specifically miniconda. You often do not need all the software that Anaconda provides. We can instead purposefully install software as and when we need it. Anaconda is also a private company. Though conda itself is free and open source, Anaconda was developed by a this private entity and has recently been pushing for [license fees](https://www.datacamp.com/blog/navigating-anaconda-licensing). It is a little unclear how this will transpire within academia as opposed to business. --- ## Installation Conda (and specifically miniconda) can be installed with from conda forge. This is an open source community that archives the many of the recipes for the packages you may want to use. https://conda-forge.org/download/ --- ## Environments The first step when using conda is to create an environment. We want to software used for different pieces of analysis to be siloed off from one another i.e. one environment for scRNAseq, ATACseq etc. We may even need to create an environment for a single piece of software if it has very specific dependencies. This is on the command line. ``` sh conda create -n demo ``` --- ## Environments If you are unsure what environments you have you can use *conda info* to find more information about all your environments. ``` sh conda info --envs ``` --- ## Activation Before you can really do anything with your software you must first activate your environment. Even if you create a new environment, it will not be used until activated. ``` sh conda activate demo ``` --- ## Install We can simply install our software using *conda install*. Here lets try installing *seaborn* a python package for plotting in python. There are many packages available. You can check on the [Anaconda search repository](https://anaconda.org/anaconda/repo). ``` sh conda install seaborn ``` --- ## Channels The conda system has several channels. These are repositories of packages that are managed by different communities. One of these is bioconda. This is a great place to get bioinformatic specific packages. If we try to install samtools from the deafult channels it cannot be found. ``` sh conda install samtools ``` By specifying the channel we can now get samtools. ``` sh conda install bioconda::samtools ``` --- ## Using Software If the environment is activated all the installed software in the environment is available to be run. ``` sh samtools help ``` --- ## Maintenance of environments As you collate and build environments you will want to make sure they are preserved and shareable. First we can check on our environment with conda list. This tells us what software we have installed and the versions. ``` sh conda list ``` --- ## Maintenance of environments Listing software versions is useful, but we want a more systematic way to preserve and detail our environments. For this we can use *conda env export*. This saves a YAML file. ``` sh conda env export > environment.yml ``` --- ## YAML ``` name: demo channels: - bioconda - conda-forge dependencies: - bzip2=1.0.8=h99b78c6_7 - c-ares=1.34.4=h5505292_0 - ca-certificates=2025.1.31=hf0a4a13_0 - htslib=1.21=hfcd771d_1 - krb5=1.21.3=h237132a_0 - libcurl=8.12.1=h73640d1_0 - libcxx=19.1.7=ha82da77_0 - libdeflate=1.22=hd74edd7_0 - libedit=3.1.20250104=pl5321hafb1f1b_0 - libev=4.33=h93a5062_2 - liblzma=5.6.4=h39f12f2_0 - libnghttp2=1.64.0=h6d7220d_0 - libssh2=1.11.1=h9cc3647_0 - libzlib=1.3.1=h8359307_2 - ncurses=6.5=h5e97a16_3 - openssl=3.4.1=h81ee809_0 - samtools=1.21=h267f7b9_1 - zstd=1.5.7=h6491c7d_1 prefix: /Users/mattpaul/Documents/RU/Software/mini/envs/demo ``` --- ## Maintenance of environments The YAML file can be viewed directly. But we can also import it using *conda env create*. Using this approach we can share the our conda envs. This *yml* file is a key way we can ensure reproducibility. ``` sh conda env create -f environment.yml ``` --- ## A few tips * Work with miniconda or even better use Herper (we will introduce this soon) * Always install software into new environments (never the base) --- ## What is Herper Herper is an R package that provides a simple toolset to install and manage Conda packages and environments from within the R console. It is built up from the package *reticulate* which is used to run python from within R. --- ## Installing Tools The **install_CondaTools()** function allows the user to specify required Conda software and the desired environment to install into. Miniconda is installed as part of the process (by default into the r-reticulate's default Conda location - /github/home/.local/share/r-miniconda) and the user's requested conda environment built within the same directory (by default /github/home/.local/share/r-miniconda/envs/USERS_ENVIRONMENT_HERE). ``` r library(Herper) install_CondaTools(tools = "star", env = "rnaseq") ``` ``` $pathToConda [1] "/Users/mattpaul/Desktop/My_Conda/bin/conda" $environment [1] "rnaseq" $pathToEnvBin [1] "/Users/mattpaul/Desktop/My_Conda/envs/rnaseq/bin" ``` --- ## Installing Tools with speicific conda If you already have Miniconda installed or you would like to install to a custom location, you can specify the path with the *pathToMiniConda* parameter. ``` r my_miniconda <- "~/Desktop/My_Conda" install_CondaTools(tools = "star", env = "rnaseq", pathToMiniConda = my_miniconda) ``` --- ## Adding more tools It's easy to add more tools. You just use *install_CondaTools()* again, but an extra argument also needs to be added: *updateEnv = TRUE*. By default when we run this command, the path to Conda and the environment is returned. We can save this as a variable for later. ``` r conda_paths <- install_CondaTools(tools = c("salmon", "kallisto"), env = "rnaseq", updateEnv = TRUE, pathToMiniConda = my_miniconda) conda_paths ``` ``` $pathToConda [1] "/Users/mattpaul/Desktop/My_Conda/bin/conda" $environment [1] "rnaseq" $pathToEnvBin [1] "/Users/mattpaul/Desktop/My_Conda/envs/rnaseq/bin" ``` --- ## Checking packages in an environment The **list_CondaPkgs** function allows users to check what packages are installed in a given environment. ``` r list_CondaPkgs("rnaseq", pathToMiniConda = my_miniconda) ``` ``` boost-cpp 1.78.0 conda-forge osx-64 bzip2 1.0.8 pkgs/main osx-64 c-ares 1.19.0 pkgs/main osx-64 ca-certificates 2023.05.30 pkgs/main osx-64 hdf5 1.12.2 conda-forge osx-64 htslib 1.17 bioconda osx-64 icu 70.1 conda-forge osx-64 kallisto 0.50.0 bioconda osx-64 krb5 1.20.1 pkgs/main osx-64 libaec 1.0.6 conda-forge osx-64 ``` --- ## Finding Conda packages If the user is unsure of the exact name, or version of a tool available on conda, they can use the **conda_search** function. Searches will find close matches for incorrect queries. ``` r conda_search("kall", pathToMiniConda = my_miniconda) ``` ``` There are no exact matches for the query 'kall', but multiple packages contain this text: - kallisto - r-merge-kallisto ``` --- ## Finding Conda package versions If you have the exact name you can search for what versions are available on Conda. ``` r conda_search("kallisto", pathToMiniConda = my_miniconda) ``` ``` 2 kallisto 0.43.1 https://conda.anaconda.org/bioconda/osx-64 4 kallisto 0.44.0 https://conda.anaconda.org/bioconda/osx-64 5 kallisto 0.45.0 https://conda.anaconda.org/bioconda/osx-64 6 kallisto 0.45.1 https://conda.anaconda.org/bioconda/osx-64 8 kallisto 0.46.0 https://conda.anaconda.org/bioconda/osx-64 9 kallisto 0.46.1 https://conda.anaconda.org/bioconda/osx-64 12 kallisto 0.46.2 https://conda.anaconda.org/bioconda/osx-64 15 kallisto 0.48.0 https://conda.anaconda.org/bioconda/osx-64 16 kallisto 0.50.0 https://conda.anaconda.org/bioconda/osx-64 ``` --- ## Finding Conda package versions Specific package versions can be searched for using the Conda format i.e. "kallisto==0.46", "kallisto>=0.48" or "kallisto<=0.45". ``` r conda_search("kallisto<=0.45", pathToMiniConda = my_miniconda) ``` ``` 2 kallisto 0.43.1 https://conda.anaconda.org/bioconda/osx-64 4 kallisto 0.44.0 https://conda.anaconda.org/bioconda/osx-64 ``` --- ## Installing specific versions We can use the same version nomenclature to also install these tools. Here we will downgrade the version of Kallisto we have installed. ``` r conda_paths <- install_CondaTools(tools = "kallisto<=0.45", env = "rnaseq", updateEnv = T, pathToMiniConda = my_miniconda) ``` --- ## Installing specific versions We can now see that we have downgraded Kallisto. ``` r library(magrittr) library(dplyr) list_CondaPkgs("rnaseq", pathToMiniConda = my_miniconda) %>% dplyr::filter(name == "kallisto") ``` --- ## Running software Once installed within a Conda environment, many external software can be executed directly from the Conda environment's bin directory without having to perform any additional actions. The Herper package uses the **[withr](https://withr.r-lib.org)** family of functions (**with_CondaEnv()** and **local_CondaEnv()**) to provide methods to **temporarily** alter the system PATH and to add or update any required environmental variables. This is done without formally activating your environment or initializing your conda. --- ## **with_CondaEnv** The **with_CondaEnv** allows users to run R code with the required PATH and environmental variables automatically set. The **with_CondaEnv** function simply requires the name of conda environment and the code to be executed within this environment. Additionally we can also the **pathToMiniconda** argument to specify any custom miniconda install location. Here we use the **with_CondaEnv** to allow us to *see* the tools installed in out our "rnaseq" environment. We can then use the R function *system2()* to run some terminal/command line code from within R. In this case we want to run the salmon help. ``` r res <- with_CondaEnv("rnaseq", system2(command="salmon",args = "help",stdout = TRUE), pathToMiniConda=my_miniconda) res ``` ``` [1] "salmon v1.10.2" [2] "" [3] "Usage: salmon -h|--help or " [4] " salmon -v|--version or " [5] " salmon -c|--cite or " [6] " salmon [--no-version-check] <COMMAND> [-h | options]" [7] "" [8] "Commands:" [9] " index : create a salmon index" [10] " quant : quantify a sample" [11] " alevin : single cell analysis" [12] " swim : perform super-secret operation" [13] " quantmerge : merge multiple quantifications into a single file" ``` --- ## **local_CondaEnv** The **local_CondaEnv** function acts in a similar fashion to the **with_CondaEnv** function: it will update the required PATH and environmental variable so you can access the tools you need. The PATH and environmental variables will persist though until the current function ends. **local_CondaEnv** is best used within a user-created function, allowing access to the Conda environment's PATH and variables from within the the function itself but resetting all environmental variables once complete. ``` r salmonHelp <- function() { local_CondaEnv("rnaseq", pathToMiniConda = my_miniconda) helpMessage <- system2(command = "salmon", args = "help", stdout = TRUE) helpMessage } salmonHelp() ``` ``` [1] "salmon v1.10.2" [2] "" [3] "Usage: salmon -h|--help or " [4] " salmon -v|--version or " [5] " salmon -c|--cite or " [6] " salmon [--no-version-check] <COMMAND> [-h | options]" [7] "" [8] "Commands:" [9] " index : create a salmon index" [10] " quant : quantify a sample" [11] " alevin : single cell analysis" [12] " swim : perform super-secret operation" [13] " quantmerge : merge multiple quantifications into a single file" ``` --- ## Reproducible environments Once you are done you have an environment that is functioning well you will want to save it. One way to back it up is to export a snapshot. For Conda this snapshot is a *.yml* file. These files contain all information about the environment you would need in order to rebuild or share it for collaboration. ``` r yml_name <- paste0("rnaseq_", format(Sys.Date(), "%Y%m%d"), ".yml") export_CondaEnv("rnaseq", yml_name, pathToMiniConda = my_miniconda) ``` --- ## Our *.yml* file The yml that is output contains all the information you need to rebuild the environment. This is also a a nice resource for when it comes to writing your methods. 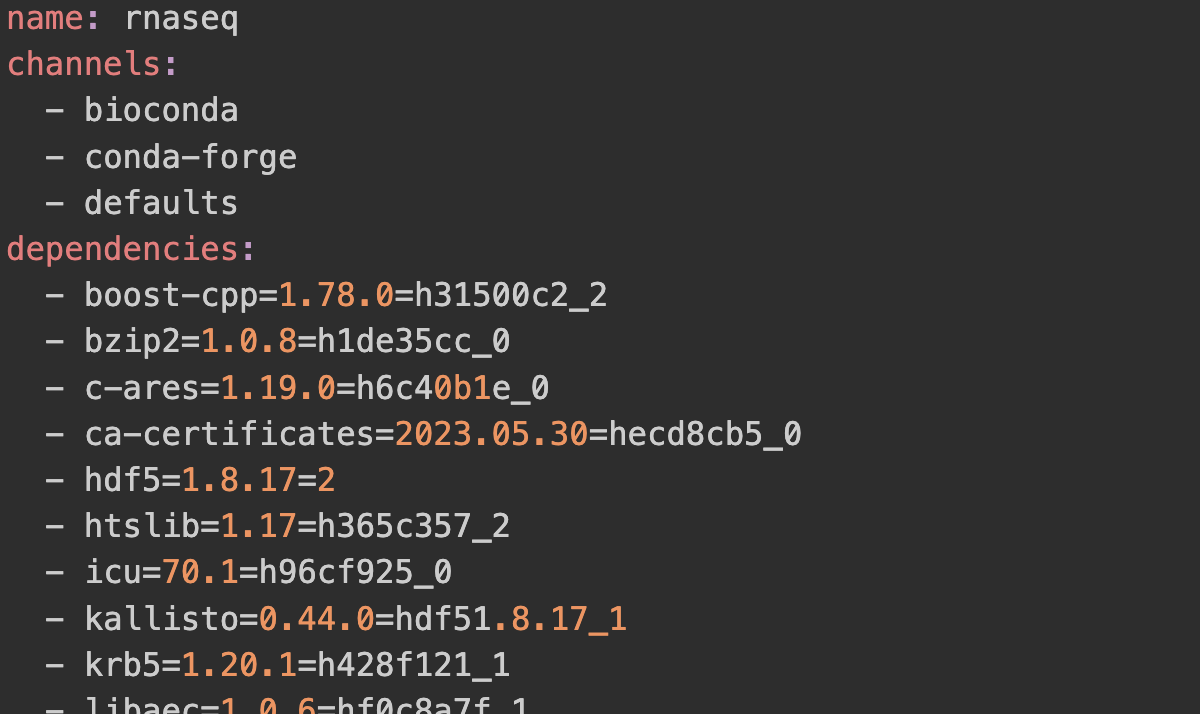 --- ## Importing from *.yml* The **import_CondaEnv** function allows the user to create a new conda environment from a *.yml* file. These can be previously exported from **export_CondaEnv**, conda, renv or manually created. Users can simply provide a path to the YAML file for import. They can also specify the environment name, but by default the name will be taken from the YAML. ``` r testYML <- system.file("extdata/test.yml", package = "Herper") import_CondaEnv(yml_import = testYML, pathToMiniConda = my_miniconda) ``` --- ## Other tricks - **Verbosity** - We can Herper either very chatty or compltely silent using the *verbose* argument. This is especially useful for troubleshooting or building pipelines. - **list_CondaEnv()** - This will list every version of Conda, and every environment you have installed. - **Mamba** - This will speed up your Conda. We are in the process of adding an option that will allow you to use Mamba resolution with Herper. --- ## What about Pip? Pip is an alternative system to install python tools. But it typically only works with python tools. Pip doesn't handle conflicting dependencies, and will upgrade/downgrade software without checking if something else depends on it. Even if this doesn't break your tools, it may give different results and hinder reproducibility as unsupported versions of tools could be used. Conda is much more considered, and checks installs to make sure that all your dependencies do not conflict. --- ## Why use Pip? * Not very Conda build is perfect * Not every tool is on Conda Sometimes we have to use pip. But that does not mean we have to leave our environment behind. --- ## Pip install First we need to mkae sure we have pip installed. Oftent his is not the case in new environments. Then we provide the direct path to our command and the isntall should run. ``` r install_CondaTools("pip", "rnaseq", pathToMiniConda = my_miniconda, updateEnv = TRUE) with_CondaEnv("rnaseq", system2(command = paste0(conda_paths$pathToEnvBin, "/pip"), args = c("install", "scanpy"), stdout = TRUE), pathToMiniConda = my_miniconda) ``` --- ## Check envs ``` r list_CondaPkgs("rnaseq", pathToMiniConda = my_miniconda) %>% dplyr::filter(name == "scanpy") ``` ``` scanpy 1.9.3 pypi pypi ``` --- ## Further Resources * The Herper [guide](https://rockefelleruniversity.github.io/Herper_Page/) * The Anaconda package [directory](https://anaconda.org/anaconda/repo) --- ## Exercises Exercise on Conda and Herper can be found [here](../../exercises/exercises/Herper_exercise.html) --- ## Contact Any suggestions, comments, edits or questions (about content or the slides themselves) please reach out to our [GitHub](https://github.com/RockefellerUniversity/Reproducible_R/issues) and raise an issue.